Hemoglobin Risk Effects (GBD 2023)
Todo
Update all instances of TODO: POST LINK with appropriate links once they are ready following the GBD 2023 update to the MNCNH portfolio simulation
Risk Overview
See the pre-print for the GBD burden of proof model of low hemoglobin publication here
GBD 2023 Modeling Strategy
Note
The low hemoglobin risk factor was NOT included in the GBD 2023 risk factors capstone publication (release ID 16). However, estimates specific to the low hemoglobin risk factor that were used to inform the low hemoglobin burden of proof publication are available within release ID 33 (see details here). Release ID 33 should be used to pull all data related to the low hemoglobin risk effects – note that release ID 16 may return data, but it is expected to be outdated.
Note that low hemoglobin risk exposure values are expected to be similar between release ID 16 and 33, although there appear to be at least slight rounding differences. We have used release ID 16 to inform low hemoglobin risk exposure in the MNCNH portfolio model to maximize consistency with GBD anemia estimates (affected by hemoglobin exposure). However, if we were to utilize the low hemoglobin PAF values directly in our model (we calculate custom PAFs for the MNCNH portfolio simulation), it may be preferable to inform hemoglobin risk exposure from release ID 33 rather than 16.
See the pre-print for the GBD burden of proof model of low hemoglobin publication here for all details on the low hemoglobin risk modeling strategy.
In GBD 2023, the hemoglobin risk effects are modeled as continuous risk curves with 1,000 exposure estimates ranging between values of 40 and 150. Exposure values <40 are assigned a risk value consistent with an exposure of 40 and exposure values >150 are assigned a risk value consistent with an exposure of 150.
Outcome |
Outcome type |
Outcome ID |
Affected measure |
Note |
|---|---|---|---|---|
Maternal hemorrhage |
cause |
367 |
incidence (affects mortality and morbidity in GBD) |
In GBD shared functions, but expected to be updated |
Maternal sepsis and other infections |
cause |
368 |
incidence (affects mortality and morbidity in GBD) |
In GBD shared functions, but expected to be updated |
Maternal hypertensive disorders |
cause |
369 |
incidence (affects mortality and morbidity in GBD) |
In GBD shared functions, but expected to be updated |
Depressive disorders |
cause |
567 |
incidence (affects mortality and morbidity in GBD) |
In GBD shared functions, but expected to be updated |
Todo
Determine how GBD handles the fact that the hemoglobin risk factor is specific to the pregnant population but the depressive disorders cause is not when we get relevant documentation
The relative risk curves for maternal disorders affected outcomes in GBD shared functions as of March 2025 are shown below for reference. These values have been transformed to be relative to a hemoglobin exposure of 110 g/L (the threshold for anemia in pregnancy) for ease of interpretation. However, they are stored in GBD shared functions as relative to a hemoglobin exposure of 40 g/L.
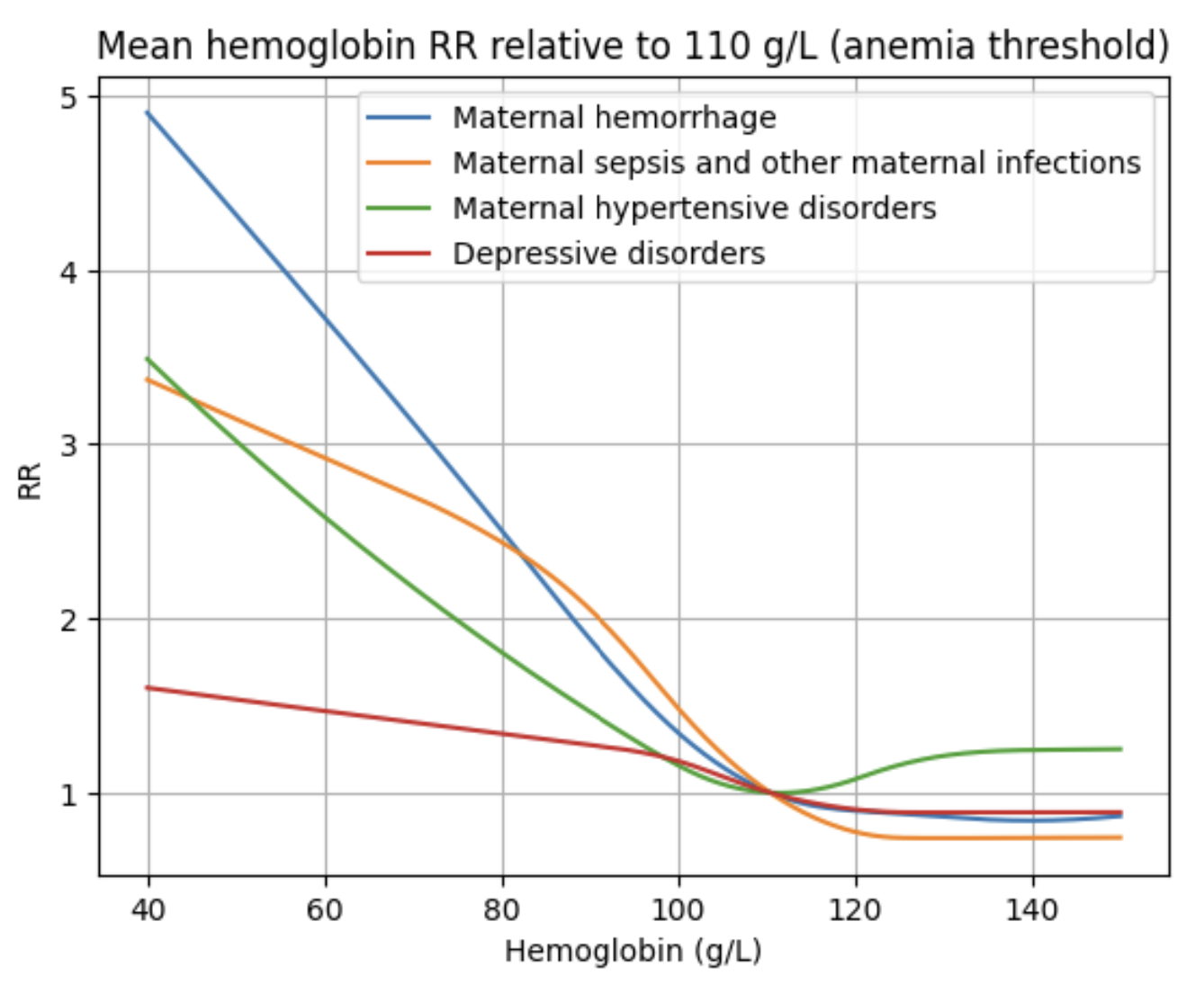
The hemoglobin team has also estimated risk effects for several additional outcomes. A list of these affected outcomes is shown below and risk curve values for each of these outcomes can be found at /mnt/team/anemia/pub/bop/sim_studies/. These risk curves were provided by the low hemoglobin risk modeller Ditha Nanditha in July of 2025.
Low birth weight (Operationalized as categorical for <2,500 grams and additional severities)
Preterm birth (Operationalized as categorical for <37 weeks and additional severities)
Small for gestational age
Large for gestational age
Neonatal sepsis and other neonatal infections
Neonatal all-cause mortality
Stillbirth
PAFs and attributable burden for hemoglobin on causes affected by LBWSG as mediated through LBWSG can be pulled using shared functions with release ID 33 and burdenator ID 393 (as tracked here). These PAFs were calculated the custom LBWSG PAF calculation and they did not consider the direct (unmediated) effect of hemoglobin on neonatal sepsis.
Risk curves for selected neonatal outcomes are shown below:
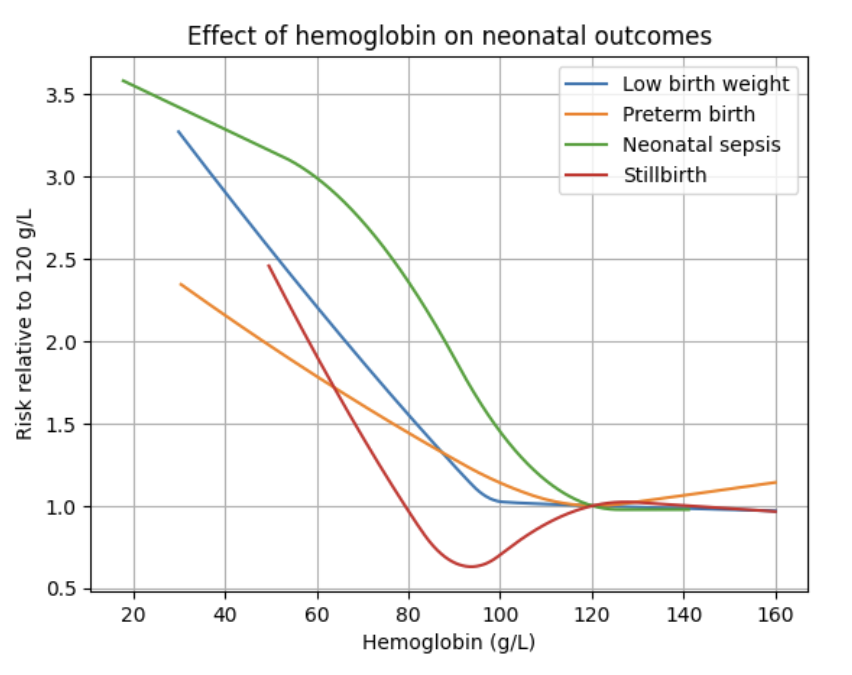
Vivarium Modeling Strategy: MNCNH Portfolio Simulation
Note that we will not be modeling direct effects of hemoglobin on the affected outcomes of stillbirth or outcomes related to gestational age or birth weight exposures. Instead, these effects will be implicitly included in the effects of the iron-containing antenatal supplementation and intravenous iron interventions, which will affect these outcomes with effect sizes that are inclusive of mediated effects through hemoglobin. We chose this modeling strategy because we obtained total effects of IFA/MMS on these outcomes from the literature and modeling direct effects from hemoglobin would require us to adjust the IFA/MMS effects for mediation through hemoglobin. So instead, we added simulated effects from IV iron on these outcomes that are calculated to be 100% mediated through hemoglobin. The modeling strategy for neonatal sepsis is different, however. This is because we do not have estimates of the total effect of either intervention on neonatal sepsis like we do for stillbirth and LBWSG outcomes. So we will model a direct effect of hemoglobin on neonatal sepsis, adjusted for mediation through LBWSG.
Category |
Outcome |
Outcome type |
Outcome ID |
Affected measure |
Note |
|---|---|---|---|---|---|
Maternal disorders |
cause |
c367 |
\(ir\) |
||
Maternal disorders |
cause |
c368 |
\(ir\) |
||
Maternal disorders |
cause |
custom |
\(ir\) |
||
Maternal disorders |
Maternal hypertensive disorders |
cause |
c369 |
TBD |
Modeling strategy for hypertensive disorders cause in the MNCNH model is still outstanding. The risk effects model for this cause may require a custom approach to account for the complexity of pre-eclampsia and related interventions in the MNCNH model. |
Neonatal outcomes |
cause |
c383 |
\(\text{CSMRisk}_{i}^\text{neonatal sepsis}\) (as opposed to \(\text{CSMRisk}_{\text{BW}_i,\text{GA}_i}^{k}\)), as defined on the MNCNH neonatal mortality page |
We will be modeling the direct effect, adjusted for mediation through LBWSG |
Todo
Update this page with hypertension cause model links when ready
Maternal disorders
Use the modeling strategy described below for the following maternal disorders subcauses:
Maternal hypertensive disorders
Todo
Link hypertension cause model documents when ready and write custom strategy for hypertensive disorders as necessary
Relative risk values to calculate custom PAFs for maternal disorders can be accessed via shared functions with the following call. Prior to the GBD 2023 update, duplicate the 250 available draws twice to obtain 500 draws such that draw 0 and draw 250 have the same value.
# return age- and cause-specific relative risk estimates for the hemoglobin risk factor
rr = get_draws(release_id=33, # GBD 2023 for topic-specific work
source='rr',
gbd_id_type='rei_id',
gbd_id=376, # hemoglobin
sex_id=2, # female for maternal disorders causes
year_id=2022, # NOTE: this call returns only one value for year ID, which is 2022. You do not need to specify a year_id, but specifying any value besides 2022 will result in a failed call
location_id=1, # NOTE: you do not need to specify a location_id. Specifying any location ID will return results specific to location_id=1
)
Before being applied in the simulation, these relative risk values are first re-scaled to be relative to the TMREL exposure value of 120 g/L. We used linear interpolation to assign RR values for exposures between points provided by the GBD risk curve. For exposure values that fall outside of the range provided by the GBD risk curve, we assign the same RR value as the nearest point on the curve. See the custom hemoglobin risk effect implementation used in the MNCNH simulation for specific details
We calculate custom PAFs for hemoglobin on maternal disorders outcome for use in the MNCNH simulation. Code to generate these PAFs can be found here TODO: POST LINK. We utilize the interactive context to initialize a population with assigned hemoglobin exposure values, assign relative risk values in accordance with observed exposure, and calculate PAFs among the population as (mean_rr - 1) / mean_rr among the age-stratified population (note that the hemoglobin risk factor data as well as maternal disorders outcome data are both already specific to sex=='female'). This PAF calculation process is limited in the following ways:
The PAF calculation does not take detailed correlation with other factors that affect maternal hemorrhage and maternal sepsis risk into account. For instance, the correlation between ANC attendance and in-facility delivery in our simulation will induce a correlation between hemoglobin (as a result of baseline IFA coverage distributed at ANC) and misoprostol coverage (available at home deliveries only), both of which affect maternal hemorrhage risk. We hypothesize that the impact of ignoring this sort of correlation will be small enough for our model to calibrate, but we can revisit this assumption if the model calibrates poorly.
We use the hemoglobin exposure distribution during pregnancy to calculate the PAF on postpartum depression rather than the hemoglobin exposure distribution among simulants who survive to the postpartum period. Given that maternal mortality is a rare event, we assume this limitation will be small, but it can be revisited if the model calibrates poorly.
Also note that the simulated population does not have a uniform distribution of maternal ages. The PAF values for age groups with less person time have less statistical precision than age groups with more person time represented in our simulation.
PAF values from our custom calculations can be found here TODO: POST LINK
Use the custom-calculated PAF values such that the maternal disorder incidence rate for an individual \(i\) for a given affected maternal disorder subcause is as follows:
Validation and Verification Criteria
Overall incidence and mortality of each affected maternal disorder cause should continue to validate to GBD
Overall hemoglobin exposure should continue to validate to GBD
Individual-level incidence and mortality of each affected maternal disorder outcome should vary according to hemoglobin exposure and corresponding risk
Note that while the mortality rate should vary in accordance with hemoglobin risk, the case fatality rate (per incident case) should not
Assumptions and Limitations
We are using population attributable fractions that do not account for correlation with confounding factors.
To minimize bias, the PAFs of hemoglobin on affected maternal disorder outcomes should account for the joint effects of any additional modeled factors that (1) are correlated with hemoglobin exposure and (2) affect the same outcome. A list of potential factors that satisfy these criteria are listed below:
Cesarean section and maternal hemorrhage
Intrapartum azithromycin intervention and maternal sepsis
Preeclampsia prevention/treatment interventions and maternal hypertensive disorders
All of these interventions may be expected to be positively correlated with hemoglobin exposure through access to the health care system or other factors. Therefore, by not considering the joint effects of these factors with the hemoglobin risk effect in baseline calibration of our model, we are likely underestimate the PAF for maternal hemorrhage and overestimate the PAF for maternal sepsis and hypertensive disorders.
This will likely result in an overestimate of the impact of interventions that work through reductions in anemia (MMS, IV iron) on maternal hemorrhage and overestimate the impact on maternal sepsis and hypertensive disorders.
We do not consider how hemoglobin exposure and/or risk effects vary with gestational age at birth in this model.
Neonatal sepsis
The effect estimates of hemoglobin on neonatal sepsis mortality as we receive them from the hemoglobin team are estimates of the total effect. However, given that there are also effects of hemoglobin on birth weight and gestational age and there is also an effect of low birth weight and short gestation (LBWSG) on neonatal sepsis, the relationship between hemoglobin and neonatal sepsis is mediated by LBWSG. We model the indirect/mediated pathway between hemoglobin and neonatal sepsis in the MNCNH portfolio model by including both the effect of hemoglobin on LBWSG (through the effects of the iron-containing antenatal supplementation and intravenous iron interventions) and the effect of LBWSG on neonatal sepsis (through the LBWSG risk effects). Therefore, when modeling the direct relationship between hemoglobin and neonatal sepsis, we must ensure that we use the direct effect estimates, which we can obtain by adjusting the total effect estimates from the hemoglobin team for mediation by LBWSG (see the risk factor mediation documentation page for background information on mediation.)
Effect derivation
Todo
Perform GBD 2023 update for effect size calculations (dependent on GBD 2023 LBWSG exposure and draw availability strategy will change)
Generally, in order to derive the direct (mediation-adjusted) effects of hemoglobin on neonatal sepsis for use in our simulation, we will use the following steps for each hemoglobin exposure level as well as each location/sex/draw pair. The code that performs these steps is hosted here, with a notebook that steps through the functions found here.
Load the estimates of the total effect of hemoglobin on neonatal sepsis (\(RR_\text{R1→O,total}\)) as obtained from the hemoglobin team. There are 250 draws available, so we duplicate them so that draw 0 has the same values as draw 250, etc.
Solve for the indirect (mediated) effect of hemoglobin on neonatal sepsis (\(RR_\text{R1→R2→O}\)). This will be done in multiple sub-steps:
Calculate the effect between hemoglobin and (separately) birth weight and gestational age at birth exposures. This will be done in a manner similar to the GBD custom calculation for the PAF of a risk on the outcome as mediated through LBWSG where we optimize for the delta value that results in the relative effect of hemoglobin on dichotomous low birth weight (LBW) and preterm birth (PTB) outcomes as estimated by the hemoglobin team. Note that these estimates are also utilized in the derivation of the effects of IV iron on GA and BW outcomes.
Calculate \(RR_\text{R1→R2→O}\) as the mean quotient of LBWSG relative risk on neonatal sepsis following application of the hemoglobin effects on birth weight and short gestation relative to the baseline value of LBWSG relative risk on neonatal sepsis. This will be done utilizing the Vivarium Interactive Context.
Calculate the direct effect (\(RR_\text{R1→O,direct}\)) as \(RR_\text{R1→O,total} / RR_\text{R1→R2→O}\) using the equation \(RR_\text{R1→O,total} = RR_\text{R1→O,direct} \times RR_\text{R1→R2→O}\).
Effects can be found in .csv files here. Each csv file contains data specific to a given draw and each file is stratified by location, sex, and hemoglobin “exposure” in grams per liter.
Data was only generated for the specific draws used in the MNCNH portfolio simulation. In generating the artifact, values for draws not used in the MNCNH portfolio simulation should ideally be filled with missing values or some illogical number like infinity so that we will notice an error if we attempt to run for a draw that does not contain valid data.
Effect application
Where,
Parameter |
Definition |
Value |
Note |
|---|---|---|---|
\(\text{CSMRisk}\) |
Neonatal sepsis cause specific mortality risk |
Defined on the neonatal sepsis cause model document |
As defined on the MNCNH neonatal mortality page, we will use \(\text{CSMRisk}_{i}^\text{neonatal sepsis}\) (as opposed to \(\text{CSMRisk}_{\text{BW}_i,\text{GA}_i}^{k}\)) |
\(\text{CSMRisk}_i\) |
Hemoglobin risk-adjusted neonatal sepsis cause-specific mortality risk for an individual simulated dyad, \(i\) |
See equation |
|
\(PAF\) |
Population attributable fraction of hemoglobin on neonatal sepsis |
Custom-calculated PAF values available here |
See details on the PAF calculation below |
\(\text{RR}_{\text{hemoglobin}_i}\) |
Relative risk value for the direct effect of hemoglobin on neonatal sepsis, specific to that individual simulated dyad’s hemoglobin exposure level at birth |
Defined in the Effect derivation section |
Notes on the PAF calculation of hemoglobin on neonatal sepsis:
The PAF of hemoglobin on neonatal sepsis is calculated according to direct (not total) effects of hemoglobin on neonatal sepsis (description of these effects found in the above section)
We do not calculate a joint PAF of hemoglobin and LBWSG on neonatal sepsis since we are not modeling correlation between hemoglobin and LBWSG in the MNCNH portfolio simulation
We use the exposure distribution of parental hemoglobin among all live and still births (not specific to parental age group) paired with the neonatal sex and age group (early neonatal and late neonatal) specific relative risk values (derivation described in the above section) for calculation of the PAF of hemoglobin on neonatal sepsis.
Code used to generate these PAF values can be found here
TODO: POST LINK
Verification and validation criteria
We should continue to meet neonatal sepsis V&V criteria in the baseline scenario
Using the interactive sim, the expected relationship between hemoglobin exposures and neonatal sepsis mortality rates unaffected by LBWSG should match the expected pattern
Assumptions and limitations
Our derivation of the effect of hemoglobin on GA and BW from estimates of PTB and LBW is limited in that it:
Assumes no change in the shape of the LBWSG exposure distribution (“shifts” are applied uniformly to the entire range of the distribution)
Does not utilize estimates of hemoglobin on additional severities of LBW or PTB
Does not consider any correlation between hemoglobin and LBWSG exposures in the derivation of the estimates of IV iron’s impact on LBWSG
We do not model correlation between hemoglobin and LBWSG and therefore do not calculate the joint PAF of these risk factors on neonatal sepsis
The PAF calculation for hemoglobin on neonatal sepsis in the late neonatal age group uses the parental hemoglobin exposure distribution among all pregnancies that result in a live or still birth rather than using the distribution among neonates who survive to the late neonatal age group. Given that hemoglobin exposure has a lower magnitude of effect on neonatal mortality than the LBWSG risk factor, we hypothesize that this is an acceptable limitation, but it can be revisited if the model calibrates poorly.